Pumas: A Pharmaceutical Modeling and Simulation toolkit
using Pumas, Plots, LinearAlgebraFor reproducibility, we will set a random seed:
using Random
Random.seed!(1)A simple one compartment oral absorption model using an analytical solution
model = @model begin
@param begin
tvcl ∈ RealDomain(lower=0)
tvv ∈ RealDomain(lower=0)
pmoncl ∈ RealDomain(lower = -0.99)
Ω ∈ PDiagDomain(2)
σ_prop ∈ RealDomain(lower=0)
end
@random begin
η ~ MvNormal(Ω)
end
@covariates wt isPM
@pre begin
CL = tvcl * (1 + pmoncl*isPM) * (wt/70)^0.75 * exp(η[1])
V = tvv * (wt/70) * exp(η[2])
end
@dynamics ImmediateAbsorptionModel
#@dynamics begin
# Central' = - (CL/V)*Central
#end
@derived begin
cp = @. 1000*(Central / V)
dv ~ @. Normal(cp, sqrt(cp^2*σ_prop))
end
endDevelop a simple dosing regimen for a subject
ev = DosageRegimen(100, time=0, addl=4, ii=24)
s1 = Subject(id=1, evs=ev, cvs=(isPM=1, wt=70))Simulate a plasma concentration time profile
param = (
tvcl = 4.0,
tvv = 70,
pmoncl = -0.7,
Ω = Diagonal([0.09,0.09]),
σ_prop = 0.04
)
obs = simobs(model, s1, param, obstimes=0:1:120)
plot(obs)Generate a population of subjects
choose_covariates() = (isPM = rand([1, 0]),
wt = rand(55:80))
pop_with_covariates = Population(map(i -> Subject(id=i, evs=ev, cvs=choose_covariates()),1:10))Simulate into the population
obs = simobs(model, pop_with_covariates, param, obstimes=0:1:120)and visualize the output
plot(obs)Let's roundtrip this simulation to test our estimation routines
simdf = DataFrame(obs)
simdf.cmt .= 1
first(simdf, 6)Read the data in to Pumas
data = read_pumas(simdf, time=:time,cvs=[:isPM, :wt])Evaluating the results of a model fit goes through an fit --> infer --> inspect --> validate cycle
julia> res = fit(model,data,param,Pumas.FOCEI())
FittedPumasModel
Successful minimization: true
Likelihood approximation: Pumas.FOCEI
Objective function value: 8363.13
Total number of observation records: 1210
Number of active observation records: 1210
Number of subjects: 10
-----------------
Estimate
-----------------
tvcl 4.7374
tvv 68.749
pmoncl -0.76408
Ω₁,₁ 0.046677
Ω₂,₂ 0.024126
σ_prop 0.041206
-----------------infer provides the model inference
julia> infer(res)
Calculating: variance-covariance matrix. Done.
FittedPumasModelInference
Successful minimization: true
Likelihood approximation: Pumas.FOCEI
Objective function value: 8363.13
Total number of observation records: 1210
Number of active observation records: 1210
Number of subjects: 10
---------------------------------------------------
Estimate RSE 95.0% C.I.
---------------------------------------------------
tvcl 4.7374 10.057 [ 3.8036 ; 5.6713 ]
tvv 68.749 5.013 [61.994 ; 75.503 ]
pmoncl -0.76408 -3.9629 [-0.82342 ; -0.70473 ]
Ω₁,₁ 0.046677 34.518 [ 0.015098 ; 0.078256]
Ω₂,₂ 0.024126 31.967 [ 0.0090104; 0.039243]
σ_prop 0.041206 3.0853 [ 0.038714 ; 0.043698]
---------------------------------------------------inspect gives you the model predictions, residuals and Empirical Bayes estimates
resout = DataFrame(inspect(res))julia> first(resout, 6)
6×13 DataFrame
│ Row │ id │ time │ isPM │ wt │ pred │ ipred │ pred_approx │ wres │ iwres │ wres_approx │ ebe_1 │ ebe_2 │ ebes_approx │
│ │ String │ Float64 │ Int64 │ Int64 │ Float64 │ Float64 │ Pumas.FOCEI │ Float64 │ Float64 │ Pumas.FOCEI │ Float64 │ Float64 │ Pumas.FOCEI │
├─────┼────────┼─────────┼───────┼───────┼─────────┼─────────┼─────────────┼──────────┼───────────┼─────────────┼───────────┼───────────┼─────────────┤
│ 1 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │
│ 2 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │
│ 3 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │
│ 4 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │
│ 5 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │
│ 6 │ 1 │ 0.0 │ 1 │ 74 │ 1344.63 │ 1679.77 │ FOCEI() │ 0.273454 │ -0.638544 │ FOCEI() │ -0.189025 │ -0.199515 │ FOCEI() │Finally validate your model with a visual predictive check
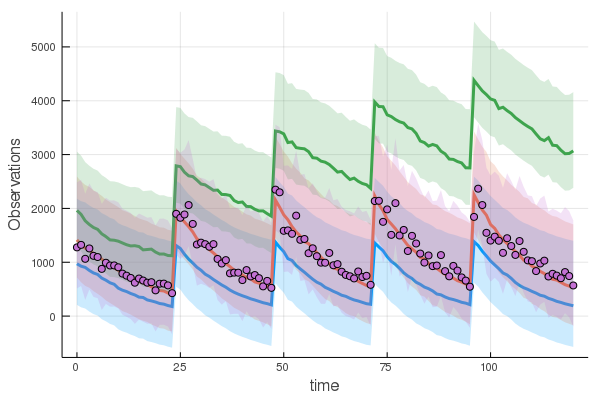
vpc(res,200) |> plotor you can do a vpc into a new design as well.
Plotting methods on model diagnostics are coming soon.
In order to simulate from a fitted model simobs can be used. The final parameters of the fitted models are available in the res.param
fitparam = res.paramYou can then pass these optimized parameters into a simobs call and pass the same dataset or simulate into a different design
ev_sd_high_dose = DosageRegimen(200, time=0, addl=4, ii=48)
s2 = Subject(id=1, evs=ev_sd_high_dose, cvs=(isPM=1, wt=70))
obs = simobs(model, s2, fitparam, obstimes=0:1:160)
plot(obs)